Logistic regression#
Logistic regression can be used to learn the probability of an event being true or false as a function of one or more features.
Here we present as example a simple ‘linear’ logistic regression.
Imports#
We import the following packages, classes, and functions.
[1]:
# for handling data:
import numpy as np
import pandas as pd
# for plotting:
import matplotlib.pyplot as plt
import halerium.core as hal
# for graphs:
from halerium.core import Graph, Entity, Variable, StaticVariable
from halerium.core.regression import linear_regression, polynomial_regression, connect_via_regression
from halerium.core.distribution import BernoulliDistribution
# for models:
from halerium.core import DataLinker, get_data_linker
from halerium.core.model import MAPModel, ForwardModel, Trainer
from halerium.core.model import get_posterior_model
# for predictions:
from halerium import Predictor
Example data#
To create example data, we simply build a forward model of logistic regression:
[2]:
n_data = 100
x_scatter = 10
with Graph("graph") as graph:
x = Variable("x", shape=(2,), mean=0, variance=x_scatter**2)
y = Variable("y", shape=(), distribution=BernoulliDistribution)
connect_via_regression(
name_prefix="parameters",
inputs=x,
outputs=y,
order=1,
)
slope = graph.parameters_y.location.slope
intercept = graph.parameters_y.location.intercept
model = ForwardModel(graph, data=DataLinker(n_data))
x_data, y_data, slope_data, intercept_data = model.get_example((x, y, slope, intercept))
data = pd.DataFrame()
data["x_1"] = x_data[:,0]
data["x_2"] = x_data[:,1]
data["y"] = y_data
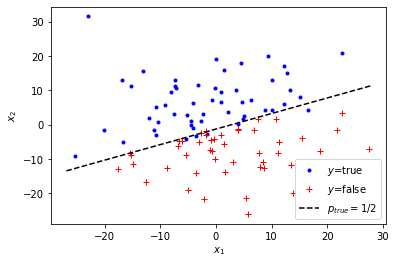
Let’s visualize the generated data:
[3]:
x_true = x_data[y_data]
x_false = x_data[~y_data]
plt.plot(x_true[:,0], x_true[:,1], '.b')
plt.plot(x_false[:,0], x_false[:,1], '+r')
r = np.linspace((-1,-1), (1, 1)) * 3 * x_scatter / np.linalg.norm(slope_data)
x_r = r * (np.array([[0,1],[-1,0]]) @ slope_data) - intercept_data / np.linalg.norm(slope_data)**2 * slope_data
plt.plot(x_r[:,0], x_r[:,1], '--k');
plt.xlabel("$x_1$");
plt.ylabel("$x_2$");
plt.legend(["$y$=true", "$y$=false", "$p_{true}=1/2$"]);
Logistic regression model#
Now let us bulid an train a logistic regression model.
[4]:
with Graph("graph") as graph:
x = Variable("x", shape=(2,), mean=0, variance=x_scatter**2)
y = Variable("y", shape=(), distribution=BernoulliDistribution)
connect_via_regression(
name_prefix="parameters",
inputs=x,
outputs=y,
order=1,
)
trained_graph = Trainer(graph=graph, data = {graph.x: data[["x_1", "x_2"]], graph.y: data["y"]})()
trained_graph.parameters_y.location.slope.mean
[4]:
<halerium.Const at 0x24cc2d0a688: name='Const', shape=(2,), global_name='graph/parameters_y/location/slope/Const', dynamic=False>
We can use the trained graph to predict the probability of y=true for a given set of x-values:
[5]:
prediction_x_data = np.linspace((-10, 0), (10, 0), 11)
predictor = Predictor(graph=trained_graph, data={trained_graph.x: prediction_x_data}, n_samples=1000)
prediction_y_data = predictor(trained_graph.y)
prediction_data = pd.DataFrame()
prediction_data["x_1"] = prediction_x_data[:,0]
prediction_data["x_2"] = prediction_x_data[:,1]
prediction_data["p_y_pred"] = prediction_y_data
display(prediction_data)
| x_1 | x_2 | p_y_pred | |
|---|---|---|---|
| 0 | -10.0 | 0.0 | 0.913 |
| 1 | -8.0 | 0.0 | 0.854 |
| 2 | -6.0 | 0.0 | 0.812 |
| 3 | -4.0 | 0.0 | 0.718 |
| 4 | -2.0 | 0.0 | 0.619 |
| 5 | 0.0 | 0.0 | 0.505 |
| 6 | 2.0 | 0.0 | 0.396 |
| 7 | 4.0 | 0.0 | 0.293 |
| 8 | 6.0 | 0.0 | 0.198 |
| 9 | 8.0 | 0.0 | 0.150 |
| 10 | 10.0 | 0.0 | 0.094 |
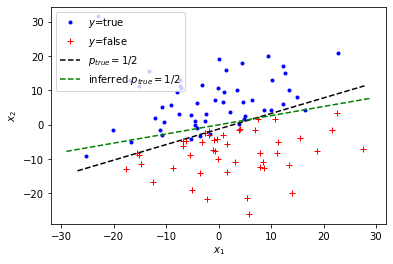
We can also compare the predicted vs. true \(p_{true}=1/2\)-line:
[6]:
x_true = data[data["y"]][["x_1", "x_2"]]
x_false = data[~data["y"]][["x_1", "x_2"]]
plt.plot(x_true["x_1"], x_true["x_2"], '.b')
plt.plot(x_false["x_1"], x_false["x_2"], '+r')
r = np.linspace((-1,-1), (1, 1)) * 3 * x_scatter / np.linalg.norm(slope_data)
x_r = r * (np.array([[0,1],[-1,0]]) @ slope_data) - intercept_data / np.linalg.norm(slope_data)**2 * slope_data
plt.plot(x_r[:,0], x_r[:,1], '--k');
inferred_slope_data = predictor(trained_graph.parameters_y.location.slope)
inferred_intercept_data = predictor(trained_graph.parameters_y.location.intercept)
s = np.linspace((-1,-1), (1, 1)) * 3 * x_scatter / np.linalg.norm(inferred_slope_data)
x_s = s * (np.array([[0,1],[-1,0]]) @ inferred_slope_data) - inferred_intercept_data / np.linalg.norm(inferred_slope_data)**2 * inferred_slope_data
plt.plot(x_s[:,0], x_s[:,1], '--g');
plt.xlabel("$x_1$");
plt.ylabel("$x_2$");
plt.legend(["$y$=true", "$y$=false", "$p_{true}=1/2$", "inferred $p_{true}=1/2$"]);
[ ]: